PestSpace molecular laboratory protocols developed
18 March 2026
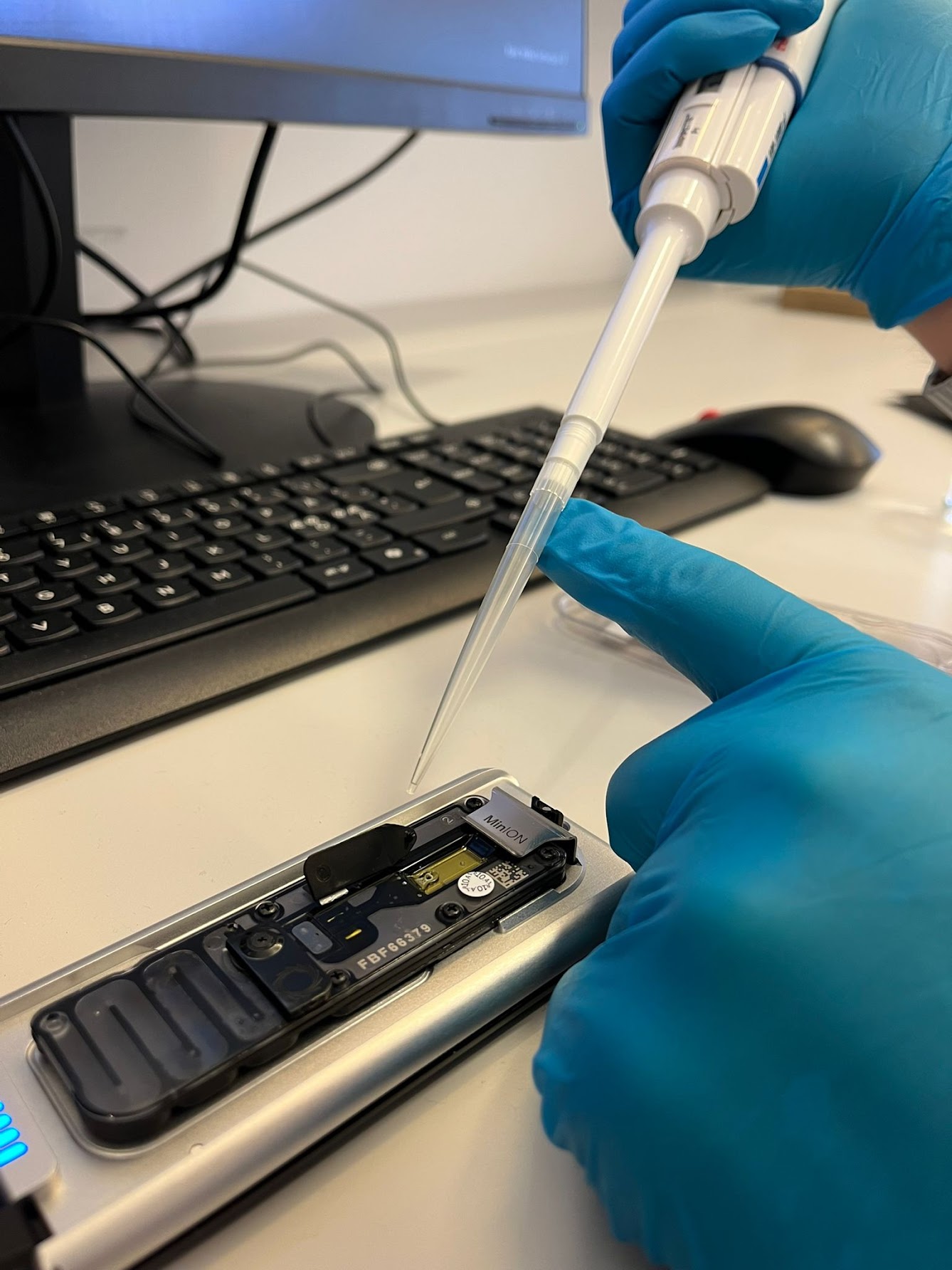
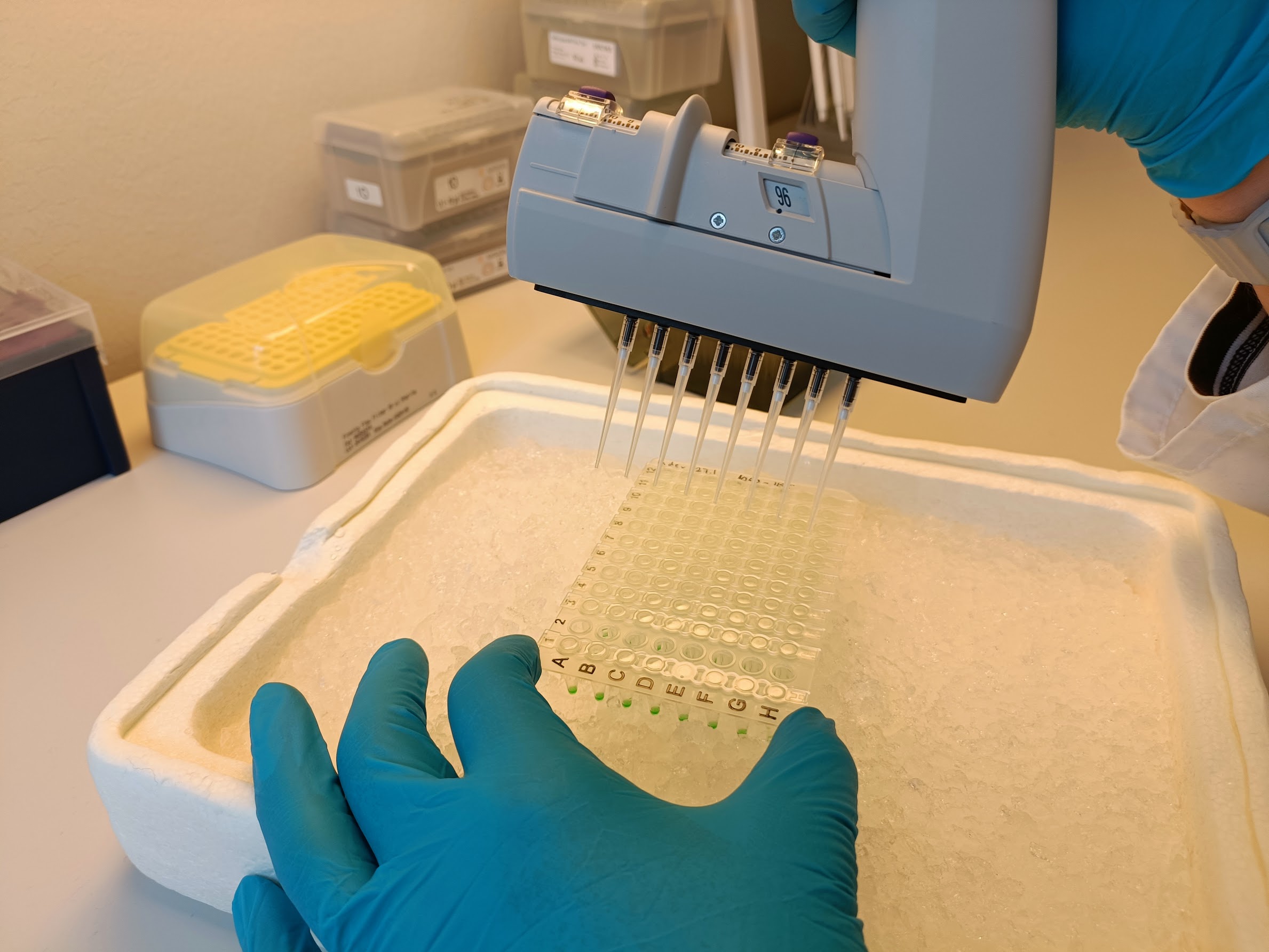
In simple terms, the process starts with extracting DNA from plant samples collected in the field. This DNA is then amplified and labelled using a two-step PCR method, and finally sequenced using a Nanopore Mk1D device. The protocol is designed to be practical, cost-effective, and scalable for larger sample sets.

As seen in the photo, Niko Johansson from the Finnish Museum of Natural History has been closely involved in developing these methods. As he explains, the process came with some challenges:
“Faba bean tends to produce a lot of plant DNA during PCR, which can interfere with the results. Using a blocking molecule helped us reduce this effect.”
DNA-based identification gives us a more precise picture of which pathogens are present in PestSpace experimental fields. It also supports the development of AI-based tools, as DNA-verified data can be used to train more accurate models in the future.
📷Take a look at a few snapshots from our research, including microscopic images.